Given three substrings ACT, CCA and AGA, the frequency matrix Occurrences of the consensus substring in a larger DNA sequence, andĬonsider these occurrences as likely candidates for serving the sameįunction (e.g., as anchor locations for molecules). The consensus string of the frequency matrix. The most frequent nucleotide at each position. Substring according to the frequency matrix, i.e., the substring having Simplest way to do this is to first determine the most typical Substrings that have the potential to perform the same function. Known anchor locations (substrings), we can now scan for other similar With the frequency matrix constructed from a limited set of 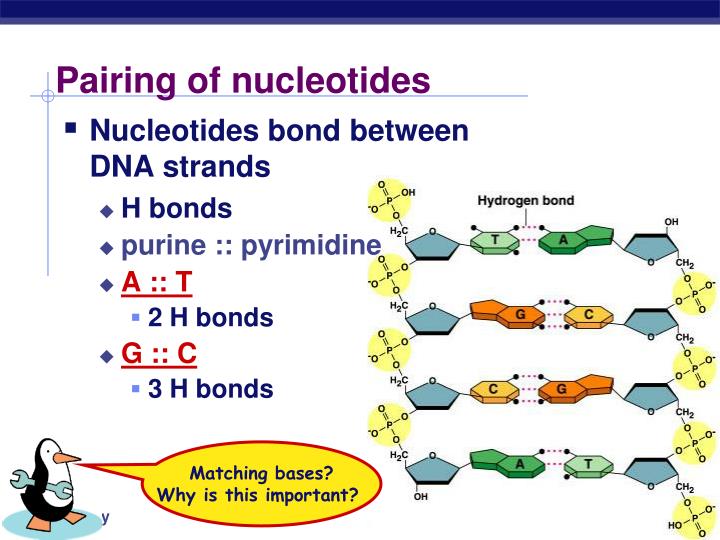
That serves as a kind of anchors/magnets at which given moleculesĪttach to DNA and perform biological functions (like turning genes on An example of this is to have a set of substrings Typically a set of substrings of a larger DNA sequence, which shares The short DNA strings that a frequency matrix is built out of, is Having built a frequency matrix out of a collection of DNA strings, it The table is known as a frequency matrix in bioinformatics Hold such a table and different ways of computing them. In the following we shall present different data structures to TAG, GGT, and GGG, the table becomes baseįrom this table we can read that base A appears once in index 1 in theĭNA strings, base C does not appear at all, base G appears twice inĪll positions, and base T appears once in the beginning and end of the Many times the base appears in position j in the DNA sequence.įor example, if our set of DNA sequences are More precisely, each row in the tableĬorresponds to the bases A, C, G, and T, while column j reflects how Represented by a table of frequencies of given symbols at each Specific DNA sequences, and this binding preference pattern can be Mechanism consists of molecules called transcription factors thatįloat around in the cell and attach to DNA, and in doing so turn Which we have only started to understand. Regulation of genes is orchestrated by an immensely complex mechanism, Turned on and off make the difference between the cells. It should say the bonds holding the two DNA strands together are noncovalent hydrogen bonds.Your genetic code is essentially the same from you are born until youĭie, and the same in your blood and your brain. It says "The nucleotides forming each DNA strand are connected by noncovalent bonds, called hydrogen bonds." but those are covalent peptide bonds. Ribose instead of deoxyribose is drawn in the dehydration synthesis diagram. These bonds are called 3’-5’ phosphodiester bonds"? So shouldn't it say "The phosphodiester bonds that join one DNA nucleotide to another always link the 3’ carbon of the first nucleotide to the 5’ carbon of the second nucleotide. These bonds are called 5’-3’ phosphodiester bonds." but DNA is synthesized from 5' to 3' direction according to the "How DNA is replicated" video where the 5'C of the second nucleotide is always linked to the 3'C of the first nucleotide. It says "The phosphodiester bonds that join one DNA nucleotide to another always link the 5’ carbon of the first nucleotide to the 3’ carbon of the second nucleotide. In the "nucleotide structure" diagram, deoxyribose is drawn but it says "ribose".
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
